They were expected to see what stuff she was made of
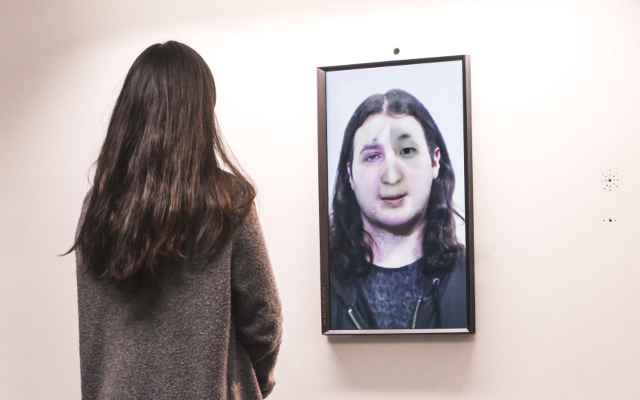
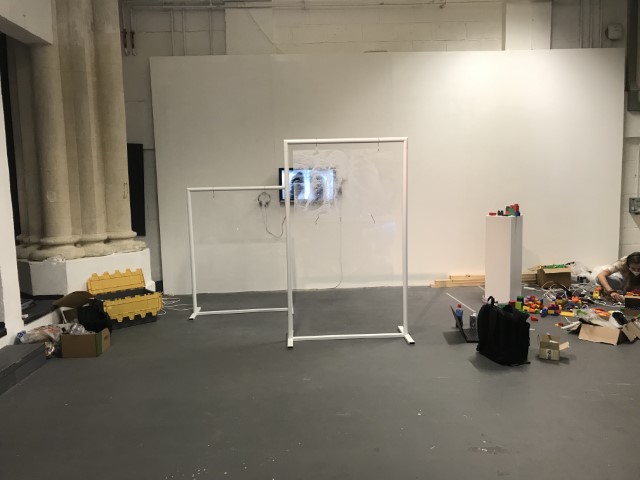
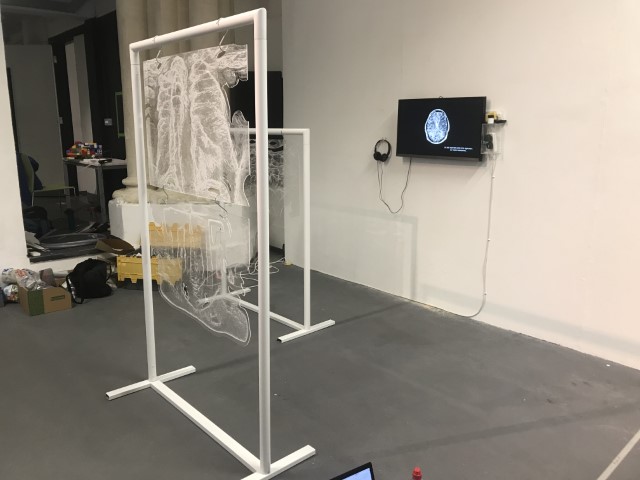
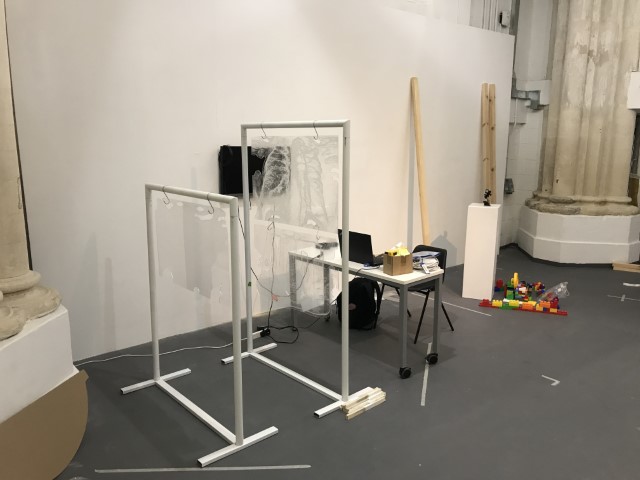
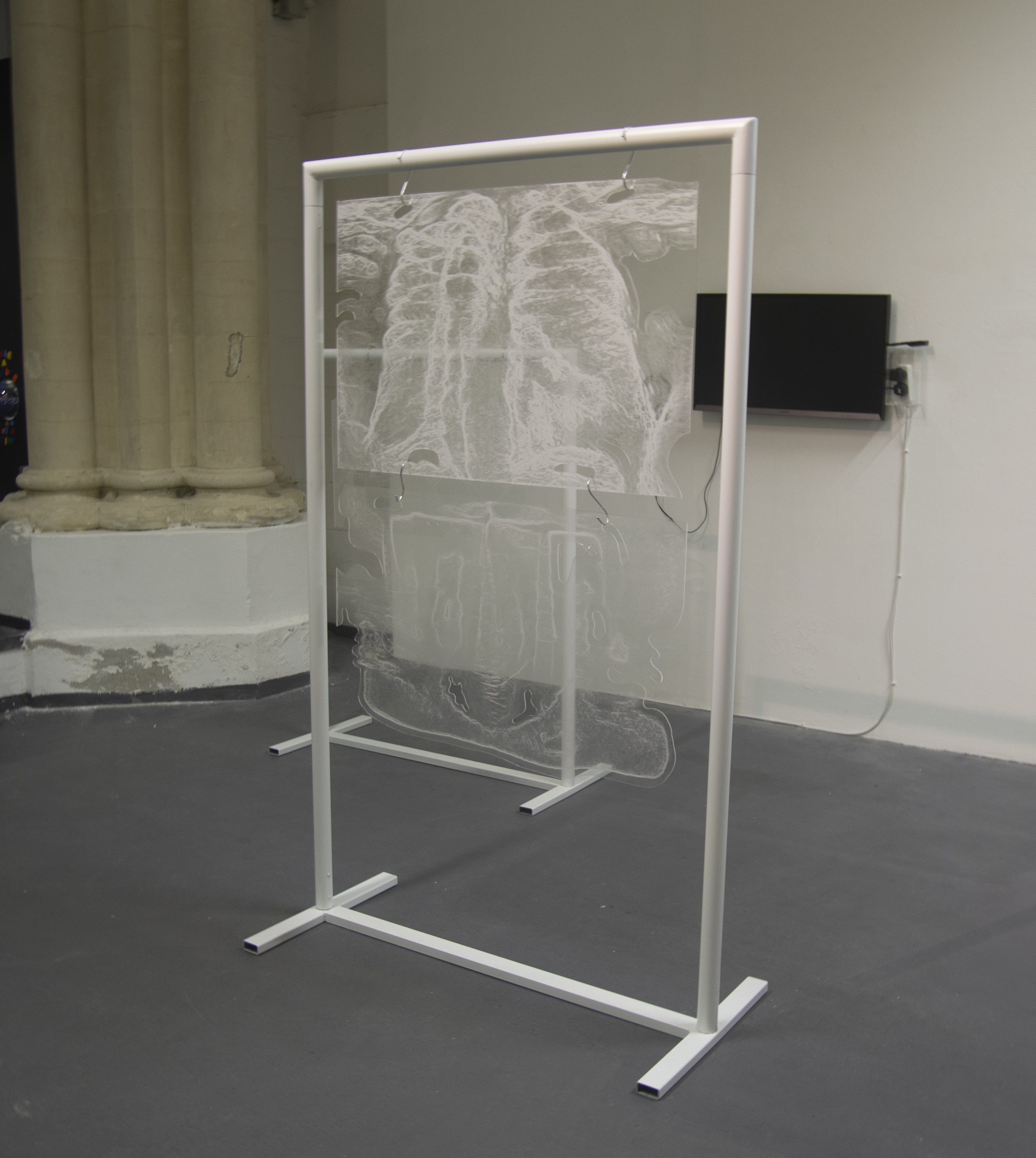
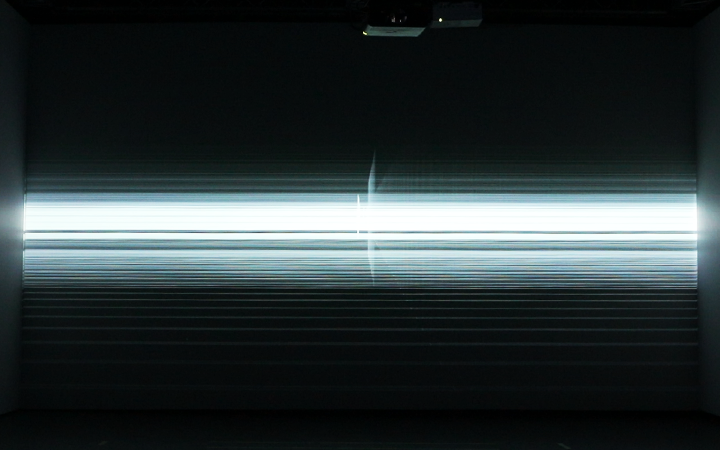
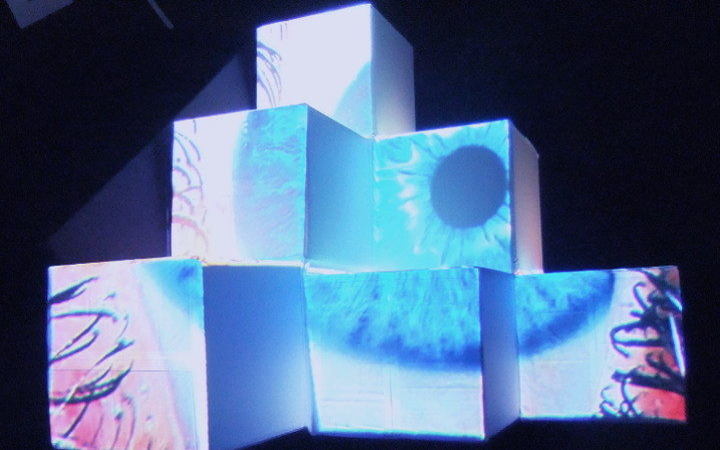
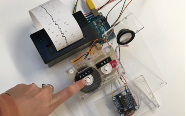
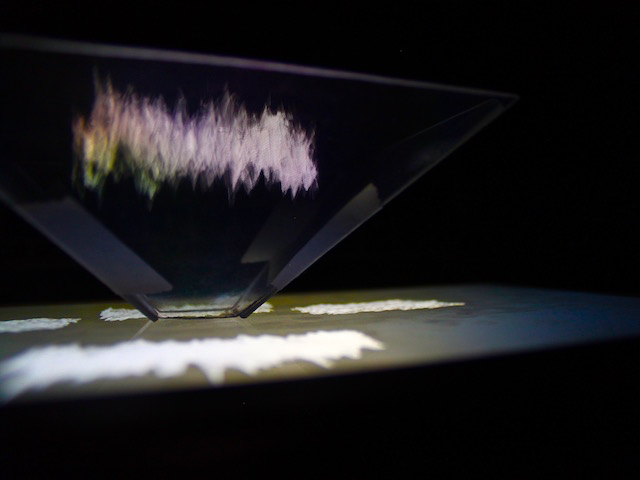
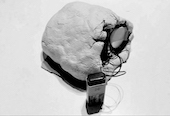
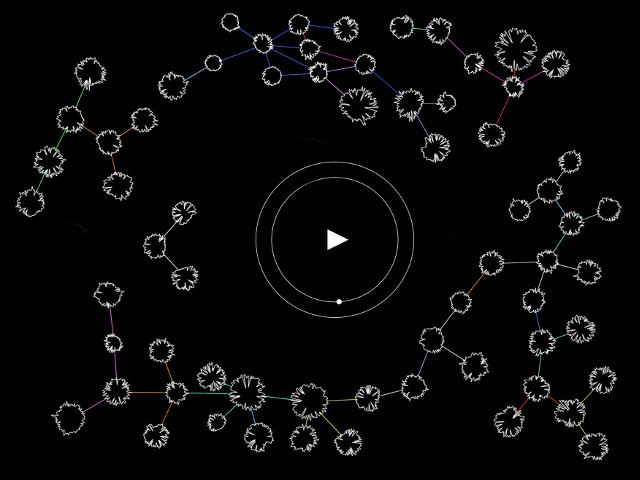
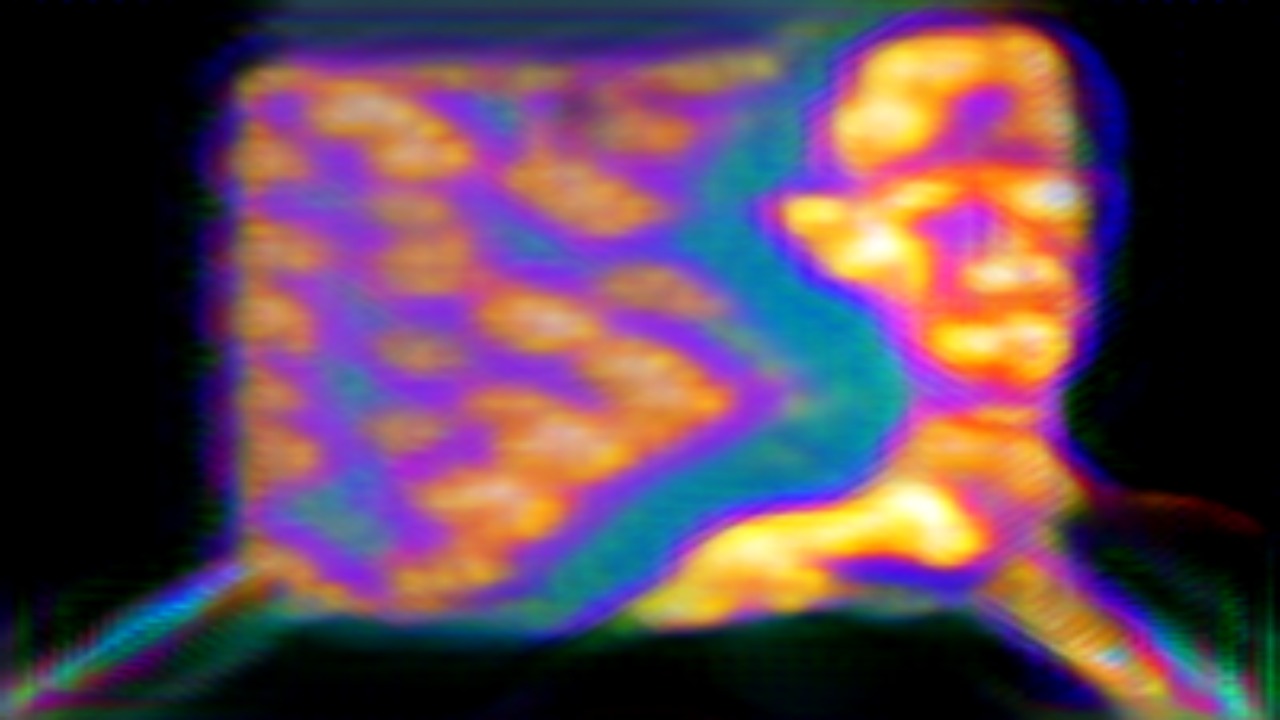
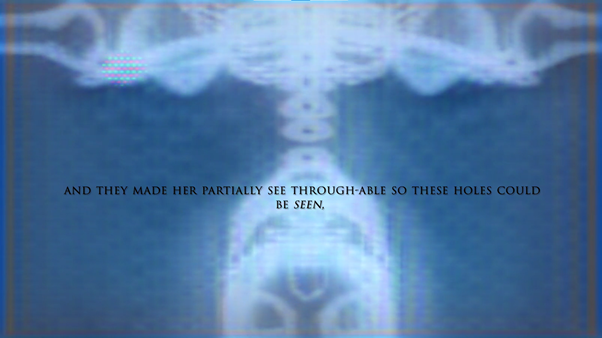
‘They were expected to see what stuff she was made of' is an audio visual installation. The focal point of the piece is a screen showing a film played on a loop. The video is overlaid with text, the tone and narrative of which draws on tropes of gothic literature, specifically themes of distrust in medical advancements and sinister scientists. The text questions the fidelity of medical imaging, specifically x-rays, PET scans and MRIs, and considers the impartiality and objectivity of the images these technologies produce and what impact the images subsequently have on the body figured via these procedures. Accompanying the screen are two free standing steel frames. Within the frames, three acrylic panels are hung with laser etched patterns that suggest anatomical illustration - one panel looks like a rib cage, another could be a pelvis or a block of flesh.
I have used several machine learning algorithms to create the video and etchings, including StyleGan2 models based on X-rays scraped from google images, pix2pix models for which I created a virtual dataset using blender and three Fast Style Transfer models trained using medical images.
produced by: Katie Tindle
Concept and background research
The video piece at the centre of the installation describes a fictionalised set of medical procedures. The film tells the story of an unknown woman who is undergoing an examination to see whether she has an illness which is not yet fully understood. The illness is causing holes and gaps to appear in the sufferers’ insides, progressing to the point where the body crumbles. Scientists are unsure of what causes the disease, whether it be environmental, moral, spiritual or caused by the procedure itself. Treatment is also debated and the full effects of the illness are not understood. This writing is the scaffolding around which the rest of the work is built, and functions as a way for me to critique the concept of a homogenous body, the language of illness and idea there can be universal truths on which to base treatment of the body. This piece acknowledges that medicine involves interpretation and to understand it to be absolute ignores the situatedness of healthcare as a practice. This denial of subjectivity in turn can work to marginalise those who have bodies which are not what is currently understood as average.
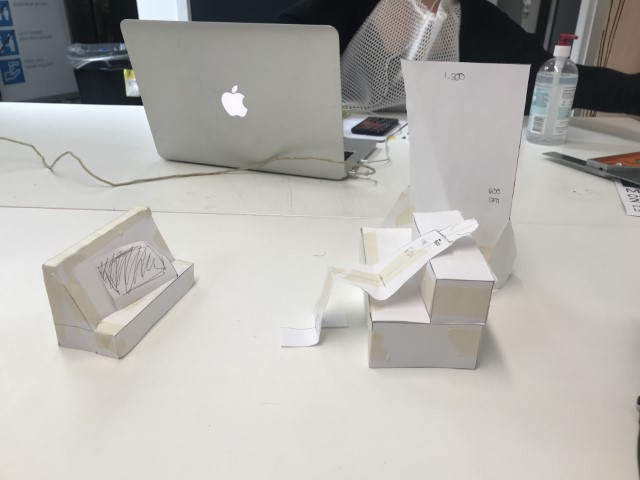
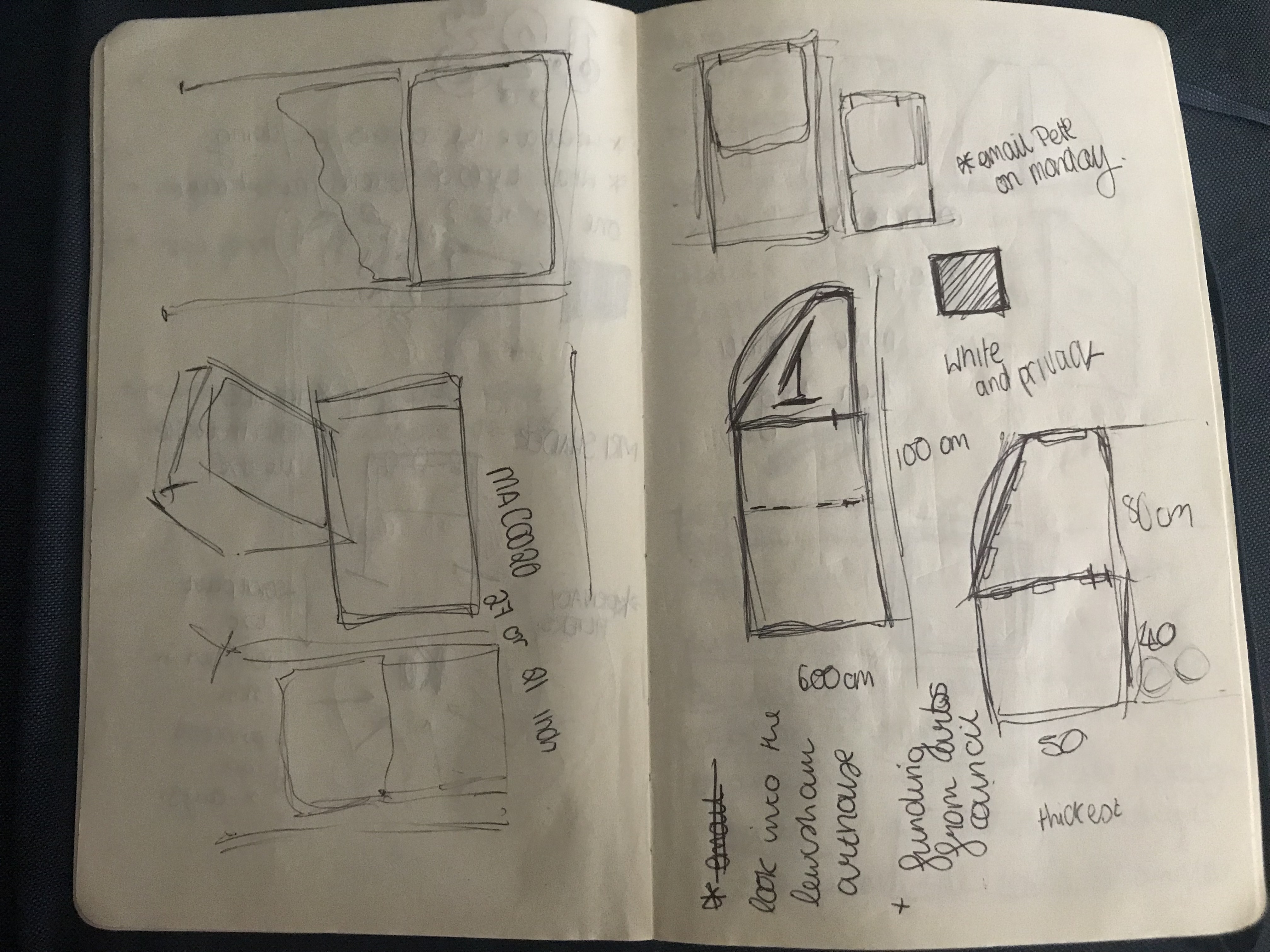
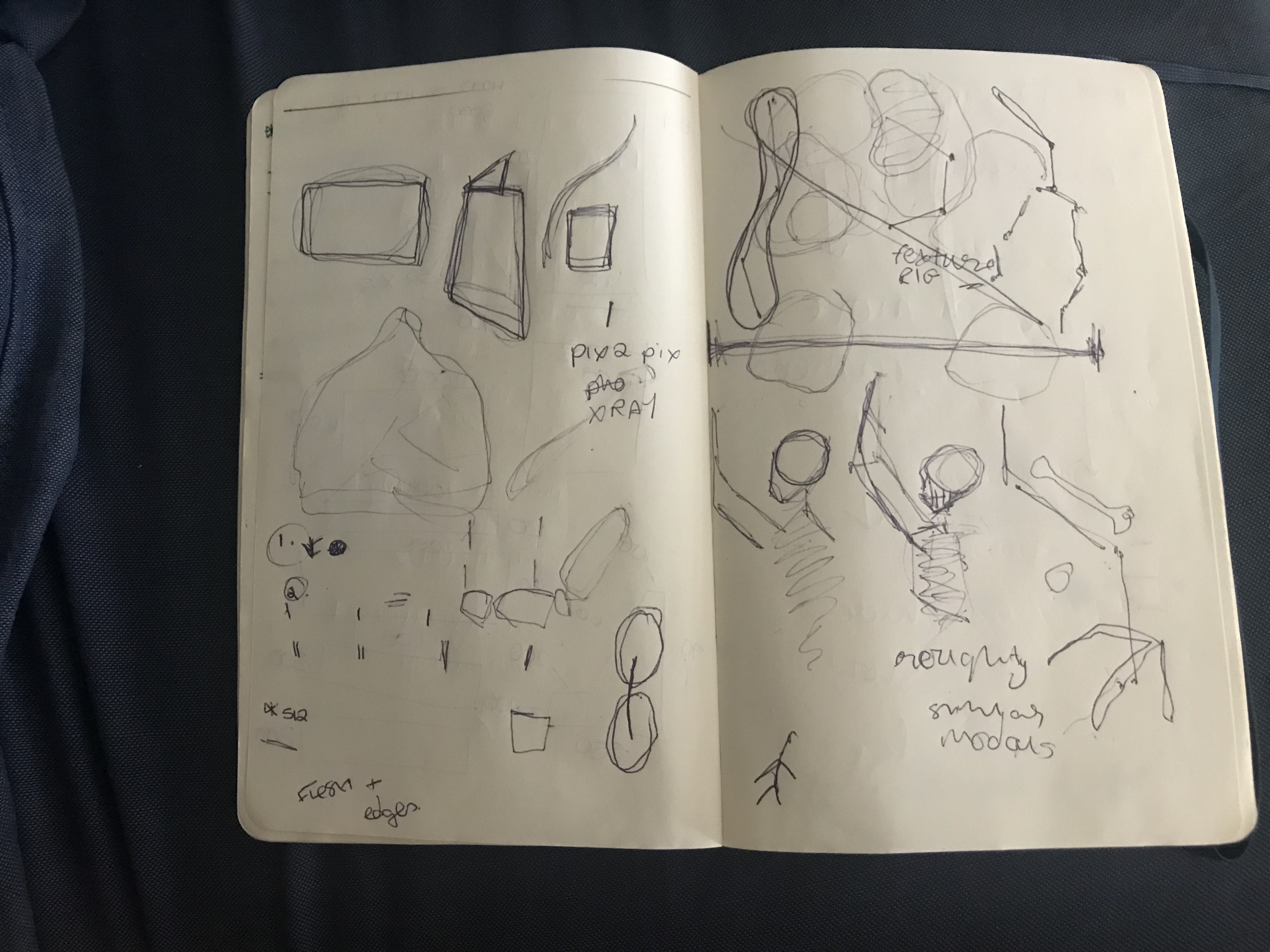
Early concept sketches.
Technical
Blob2bone, Bone2blob
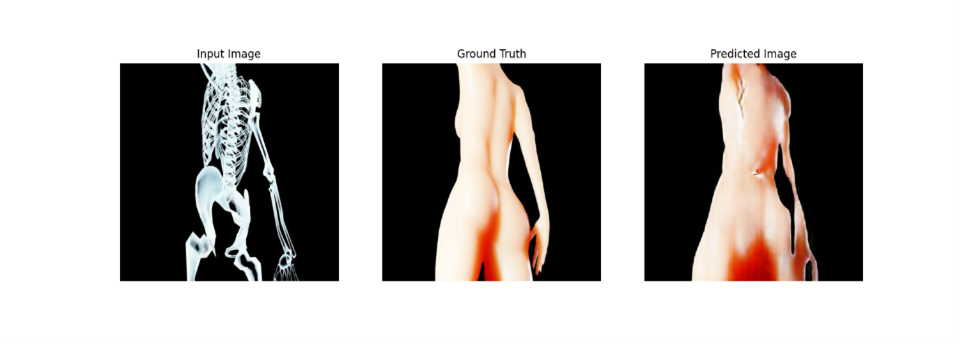
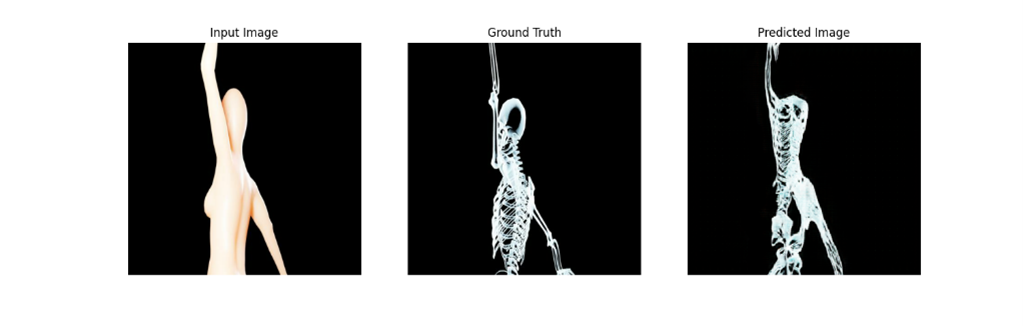
Two recurring visual motifs in the piece are made with two mirrored pix2pix models Blob2Bone and Bone2Blob. Pix2Pix is a conditional generative adversarial network (cGAN) that maps an input image to an output image. Blob2Bone takes an input image and produces an approximate skeleton based on that image. Bone2Blob takes an image of a skeleton as an input and tried to add flesh to that skeleton.
Stills of Bone2Blob and Blob2Bone taken from the final exhibition video.
Creating the Blender data set
During the development of my project I ran into an issue when looking for suitable datasets. There are many datasets available from sites like Kaggle which include thousands of medical scans, but few include a ‘before’ counterpart image. My aim was to create a pix2pix model which would generate a skeletal structure for objects, and a model which would apply flesh to a skeleton. To achieve this, I would need an A image of a person or object, and a B image of its skeleton with which to train the model. The solution was to create a virtual dataset with which augment existing material. With this in mind, I set about creating my own set of images with which to train the model using the 3D modeling software Blender.
Bone2blob during training.
Blob2bone during training.
Blender pipeline
The first thing I had to do was to find two 3D models for each half of my dataset. I found two free assets, one of a woman and one of a skeleton. I then used Blender’s shader editor to make two materials – one white skin tone, and a glowing blue semi transparent material based on the appearance of X-rays.
These are two very narrow representations of the body in many ways. Although I had considered and explored building a data set on multiple representations of bodies. However, for this piece to work as a direct comment on the homogeneisation of the body, as viewed from the perspective of machine learning in medicine, it seemed appropriate to build the model on thousands of images of the same artificial body.
Once given a material, the objects were rigged and posed in a Blender scene in the same position and manner. A camera was added to the scene which would allow me to render an image of the scene to as a jpeg. A script made each object visible in turn to the camera, which rendered its view of each object as a jpeg image. The script then moved the camera to another position to repeat the process. In this way, I was able to automate the production of thousands of paired images with which to train my model. Once the paired images were rendered, each pair were resized and concatenated into a single image using PIL. Once the data set was complete, I used it to train the pix2pix model on a Google Colab using Tensorflow.
Fast Style Transfer models
I used the above rendering method to create animations of my 3D objects for use in the video. I used a script to move the camera and render its view in tiny increments. These images were then stitched together using FFMPEG to create swooping and panning sequences, essentially stop motion animation. These sequences were then processed to make them resemble other medical imaging technologies. To achieve this, I trained Fast Style Transfer models in TensorFlow and converted the model for ML5.js. The model was then applied to the images using a p5.js programme which used ml5.js with the ml5.styleTransfer() method.
I trained style models on images of PET scans, X-Rays and MRIs scraped from google images.
Edges2Brains - pix2pix using ML5
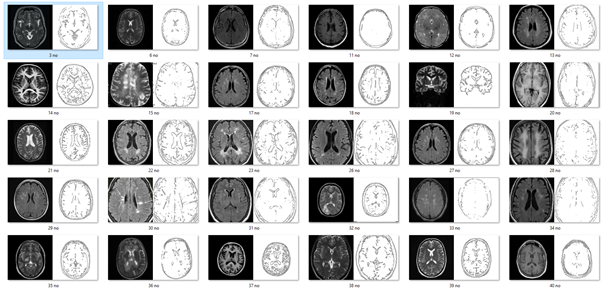
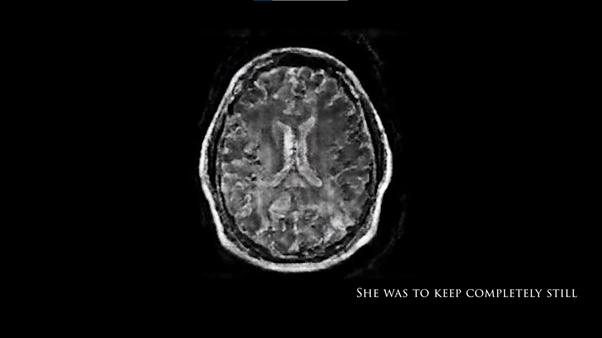
As well as the pix2pix model described above, I also created a model which generated brain MRIs based on canny edge detection. I created a data set using thousands of MRI scans paired with the edges of those images. This was trained using TensorFlow and converted the model for ML5.js. The model was then applied to the images using a p5.js programme which used ml5.js with the ml5.pix2pix() method.
When animating the brain scan sequence, I used Python Imaging Library (PIL) to interpolate between several of the edge images. I then applied the results using the p5.js programme, described above, and stitched the outcomes together using the python library FFMpeg.
Edges2Brains paired image dataset.
Edges2Brain model used in final exhibition video.
StyleGan2
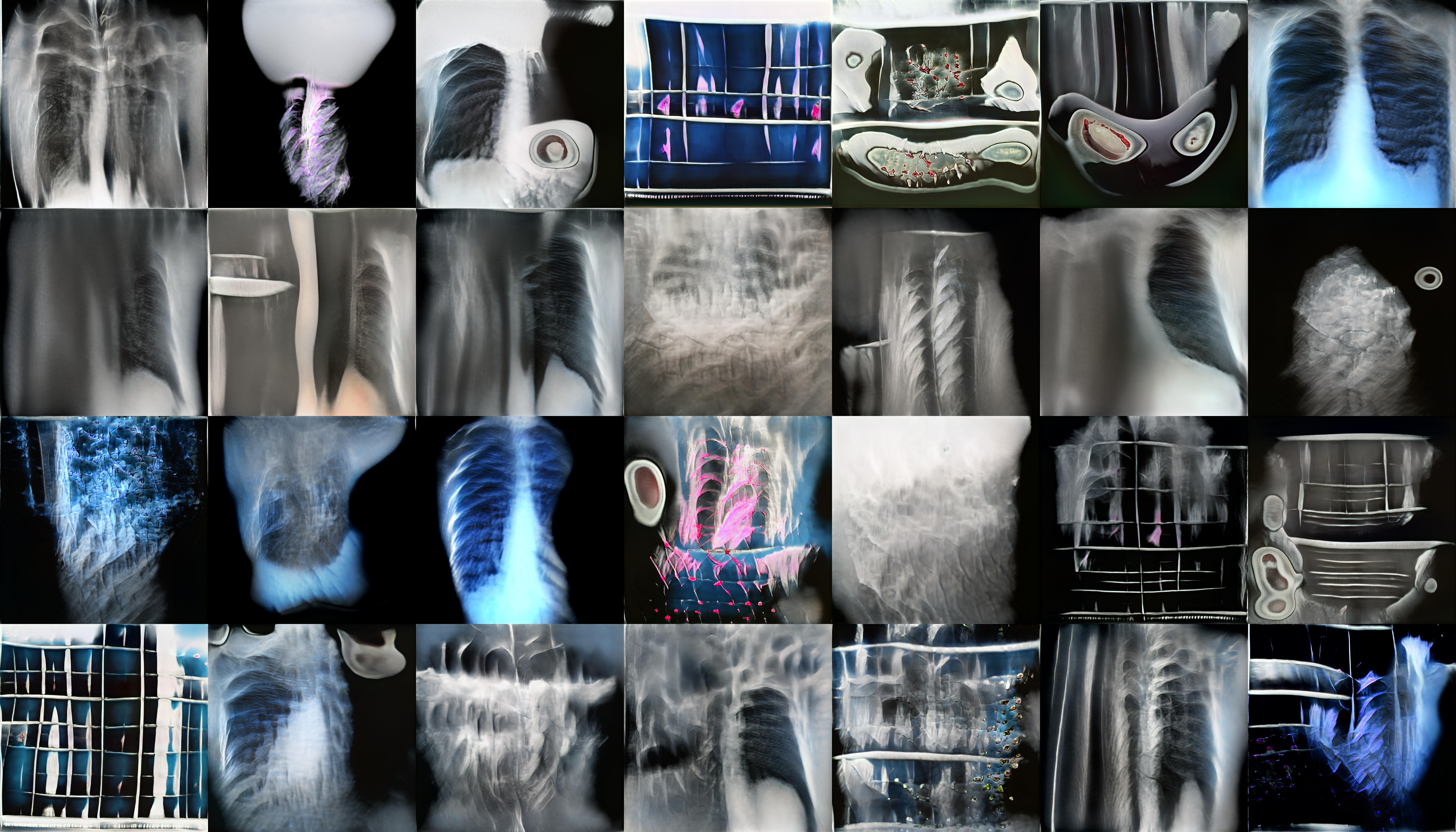
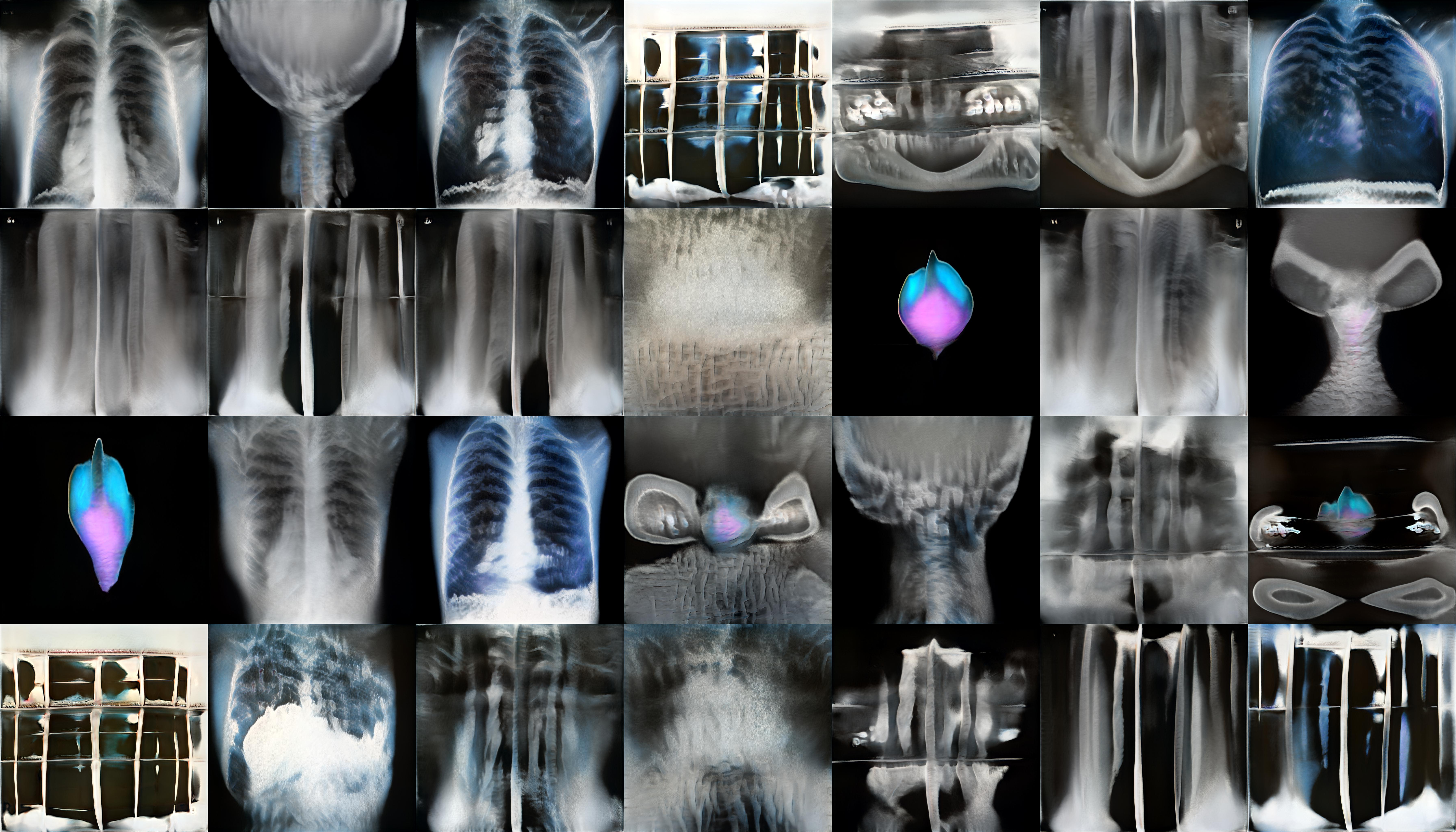
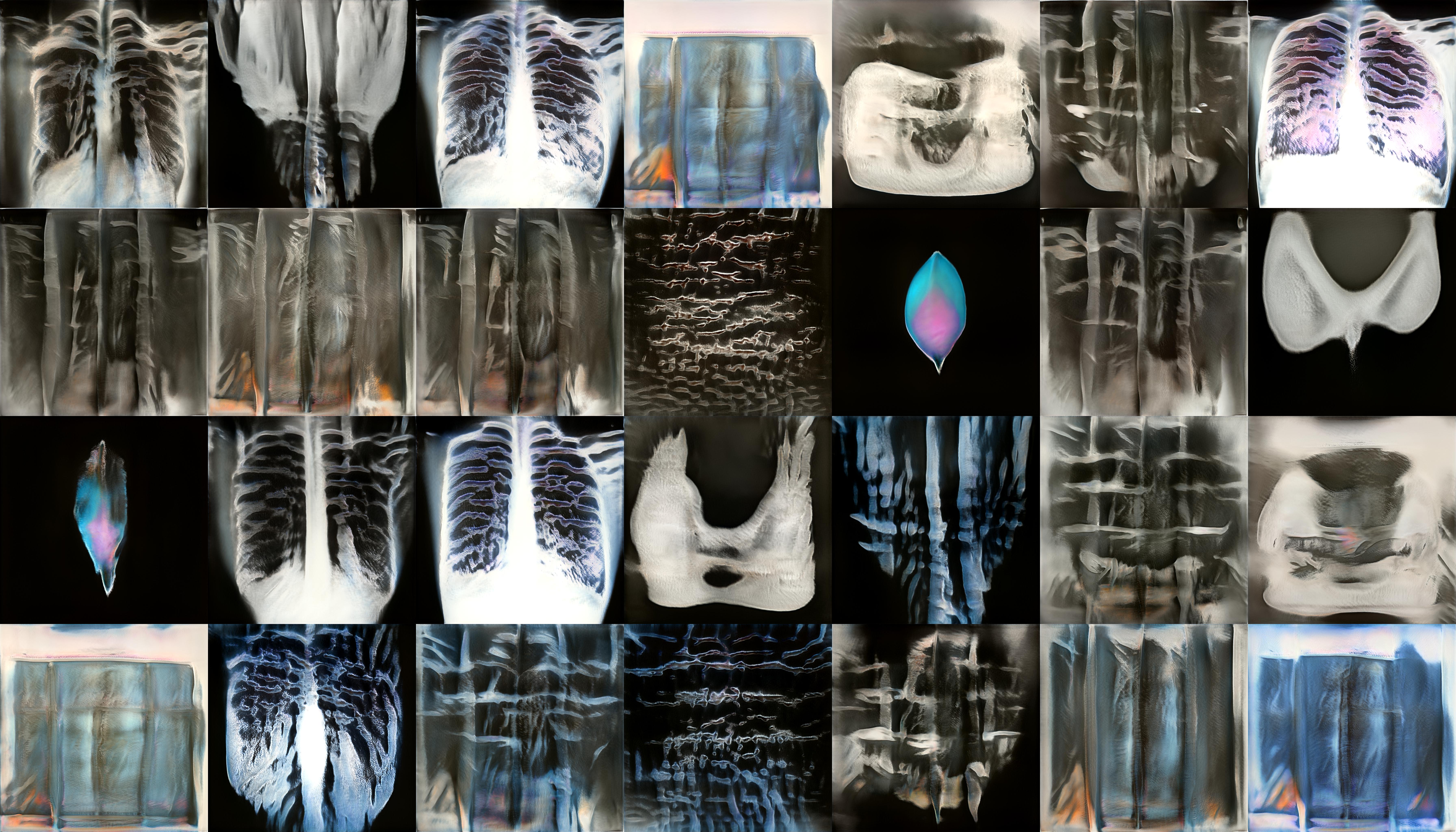
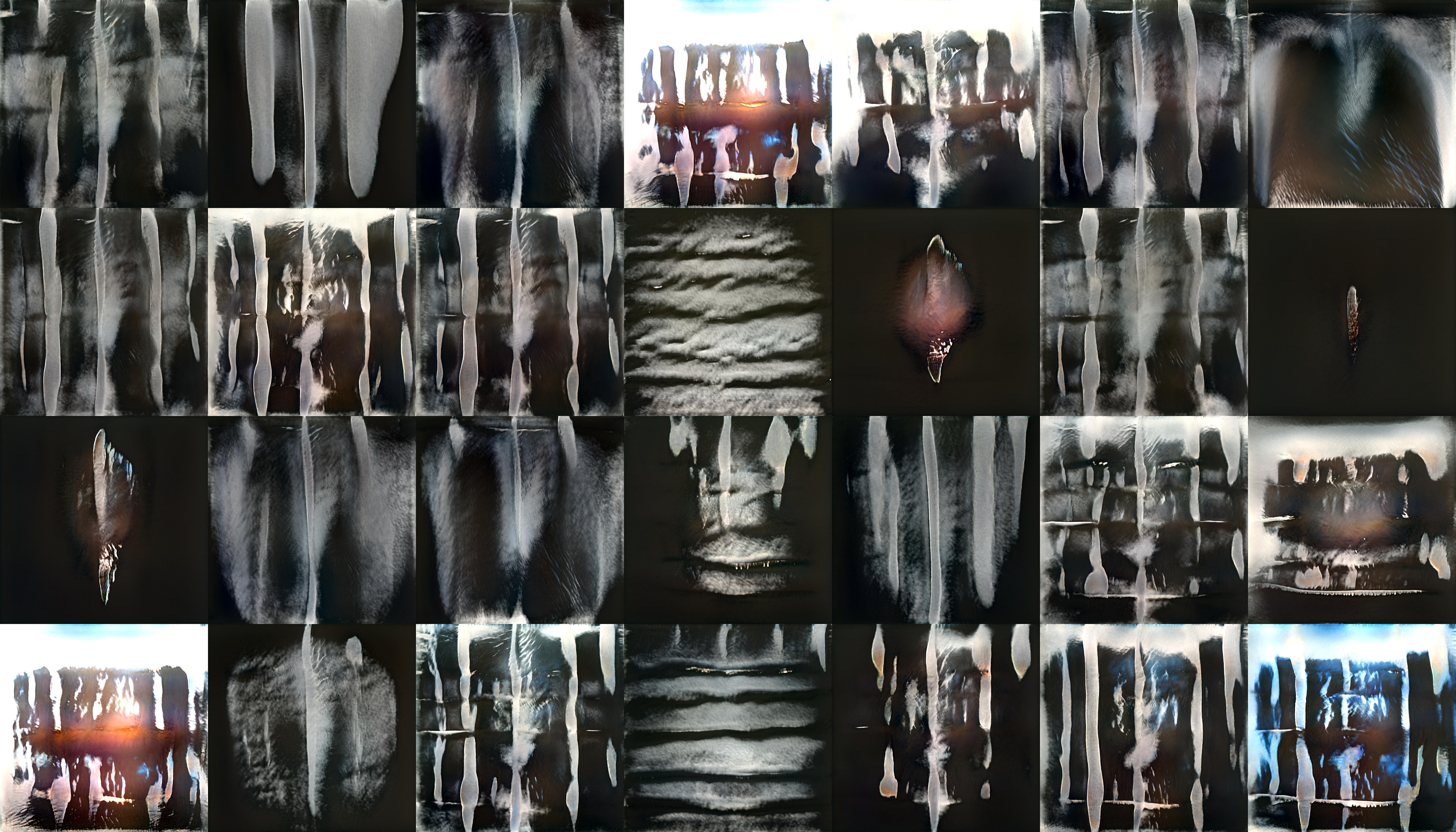
StyleGan2 is a Generative Adversarial Network which is capable of generating large quantities of high-quality images. To create the ‘realistic’ x-ray images featured in my video piece, and on the laser etched panels, I trained StyleGan2 on a set of X-rays I scraped from Google images. I used this relatively small and varied data set rather than a large medical data set as this would eventually generate weirder, more uncanny results. The majority of my time here was spent adjusting the parameters of the training and observing the training outputs.
StyleGan2 outputs during training
Trail and error
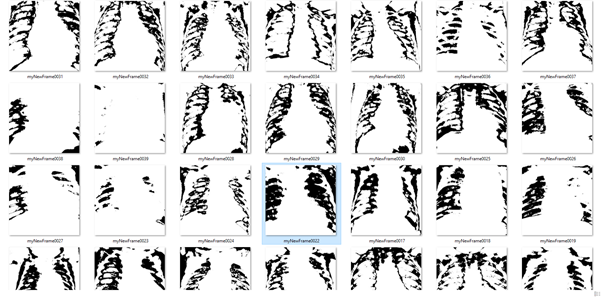
The section above describes successful techniques but there were a lot of attempted which didn’t go so well. For instance, I tried to build a set of lungs to be used an input for a pix2pix model using pixel differencing. Unfortunately this produced results that were not able to capture the complexity of the original images.
lungs with processed using pixel differencing.
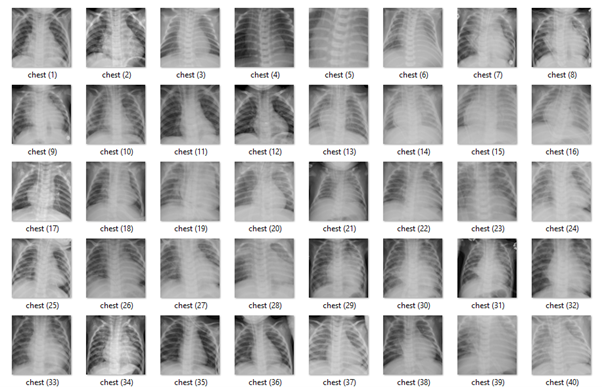
Original chest x-rays.
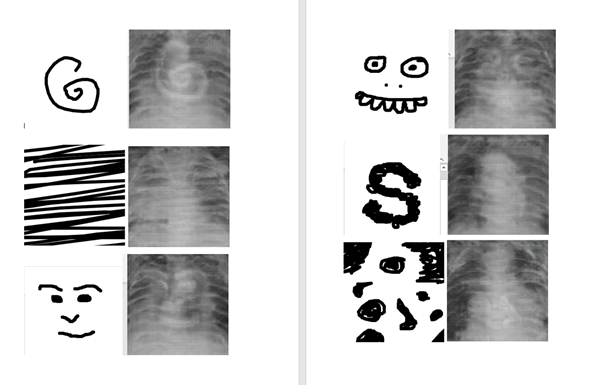
I tried again using canny edge detection, and even trained the model. The hope was that I might be able to insert drawings into the rib cages, but the results were not clear enough to be usable.
Testing different mark making with edges2lungs model.
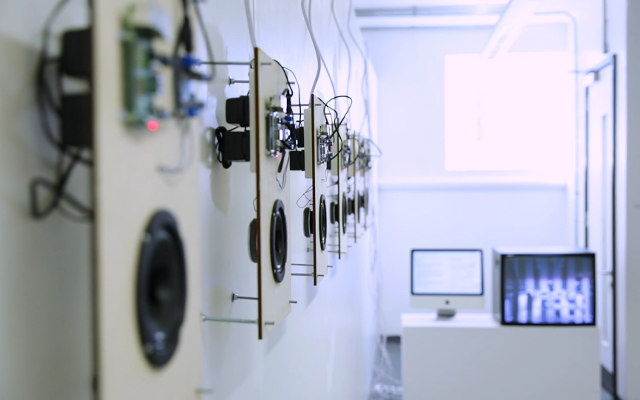
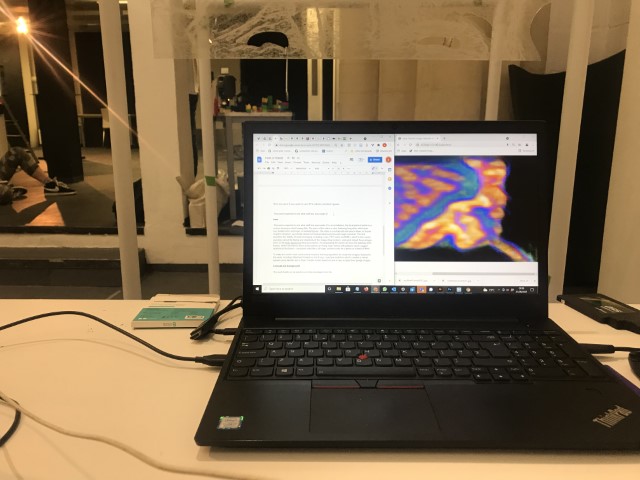
Installation
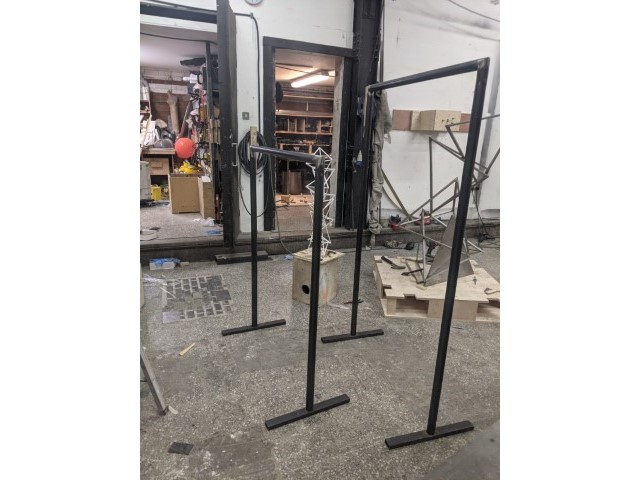
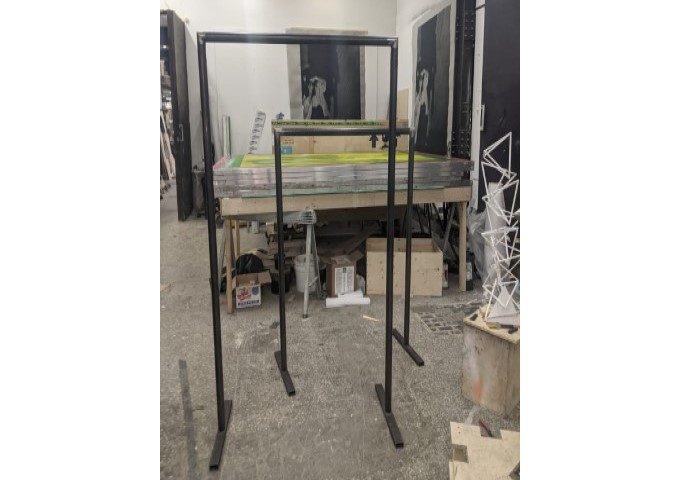
The installation was designed to seem cold and sterile. I hung the etched panels to mimic the way x-rays are hung using hangers when examined. I went through several design iterations before simplifying the set up. The screens were designed as a kind of barrier between the audience and the piece, as a lens is a barrier between a camera and its subject.
Installation sketches, development and final install.
Reflections
I intend to continue using machine learning as part of my practice as an artist. I plan to improve the blob2bone and bone2blob models, working on different scenes and lighting in blender, perfecting the poses and making models themselves more photorealistic. I would also consider restaging this piece so that the lighting better enables the etchings to be seen.
Overall, I think the work is successful. I started this project intending to make a work which built on my interest in gothic horror and existing writing practice. I hoped the work would encourage the viewer to think critically about medical imaging technologies, but avoid being didactic. I think I have achieved these goals while having pushed myself as an artist and creative technologist. I have a much greater understanding of machine learning techniques for artistic practice, and have developed new skills – including shading and scripting in blender of which I had no prior experience.
References
Brain_Tumor_Detection_MRI (no date). Available at: https://kaggle.com/abhranta/brain-tumor-detection-mri (Accessed: 25 April 2021).
Chest X-ray Images (no date). Available at: https://kaggle.com/tolgadincer/labeled-chest-xray-images (Accessed: 25 April 2021).
Duhamel, D. (2015) ‘Body Horror’, Columbia: A Journal of Literature and Art, (53), pp. 45–45.
Engstrom, L. (2016) Fast Style Transfer. Available at: https://github.com/lengstrom/fast-style-transfer (Accessed: 10 September 2021).
Mesh2Matter/blenderflow at master · georgie-png/Mesh2Matter (no date) GitHub. Available at: https://github.com/georgie-png/Mesh2Matter (Accessed: 10 September 2021).
Neural (no date) ‘Codependent Algorithms, highly calculated misunderstandings | Neural’. Available at: http://neural.it/2021/04/codependent-algorithms-highly-calculated-misunderstandings/ (Accessed: 6 May 2021).
pix2pix: Image-to-image translation with a conditional GAN (no date) TensorFlow. Available at: https://www.tensorflow.org/tutorials/generative/pix2pix (Accessed: 10 September 2021).
pix2pix-tensorflow (2021). affinelayer. Available at: https://github.com/affinelayer/pix2pix-tensorflow (Accessed: 10 September 2021).
Rapoport, M. (2020) ‘Frankenstein’s Daughters: on the Rising Trend of Women’s Body Horror in Contemporary Fiction’, Publishing Research Quarterly, 36(4), pp. 619–633. doi:10.1007/s12109-020-09761-x.
Ronay, M. and Relyea, L. (2020) ‘Body Horror’, Artforum International. Available at: http://search.proquest.com/docview/2426194575/abstract/4D06D045985845B5PQ/1 (Accessed: 5 May 2021).
says, H. tobuild a deep learning image dataset-PyImageSearch (2017) ‘How to create a deep learning dataset using Google Images’, PyImageSearch, 4 December. Available at: https://www.pyimagesearch.com/2017/12/04/how-to-create-a-deep-learning-dataset-using-google-images/ (Accessed: 10 September 2021).
Shi, Y. (2021) pix2pix in tensorflow.js. Available at: https://github.com/yining1023/pix2pix_tensorflowjs (Accessed: 10 September 2021).
Sontag, S. (1988) Illness as metaphor. 1. paperb. ed. New York, NY: Farrar, Straus and Giroux.
Steven Scott (2020) Blender Tutorial X-ray Shader ‘Cycles’. Available at: https://www.youtube.com/watch?v=W1ymJooiYJQ (Accessed: 10 September 2021).
Style Transfer training and using the model in ml5js (2021). ml5. Available at: https://github.com/ml5js/training-styletransfer (Accessed: 10 September 2021).
THE LUWIZ ART (2019) Shading a Human Skin in Blender 2.80 (Quick Tip). Available at: https://www.youtube.com/watch?v=0ybRLcBU564 (Accessed: 10 September 2021).
Writing Academic Texts Differently: Intersectional Feminist Methodolog (no date). Available at: https://www.routledge.com/Writing-Academic-Texts-Differently-Intersectional-Feminist-Methodologies/Lykke/p/book/9781138283114 (Accessed: 6 May 2021).